Protein function is decided in part by it's primary structure (the amino acids which make it up) but also it's 3D folded shape it conforms to within the body. Proteins are large polymers located within the body, with a large array of functions (from muscle to digesting food). The worker threads will send back progress while running, and the interface will automatically display these. Once the connection is valid, you can select "Generate" and begin to watch the protein fold. Otherwise specific the server here such as "". "localhost" means the gui and coordinate are run on the same machine. At this point, you can run the Graphical User Interface from any computer and select Connection. The coordinator should respond immediately saying "Got a worker". 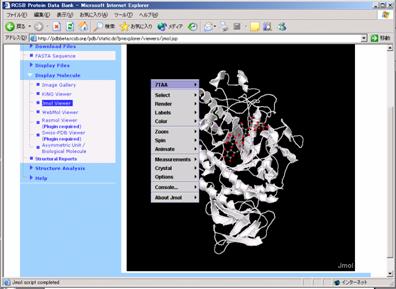
A Worker can than be started on the same machine (a worker can be run from a seperate machine). First run the coordinator and you should see "Waiting for a new connection". All three of these threads must be running in order to fold a protein. The graphical user interface (Gui.java), the worker thread which folds the protein (Worker.java), and a communication thread (Coordinator.java). Make sure all of the jars in lib are actually imported into the build path. Java3D should be installed on the machine. 
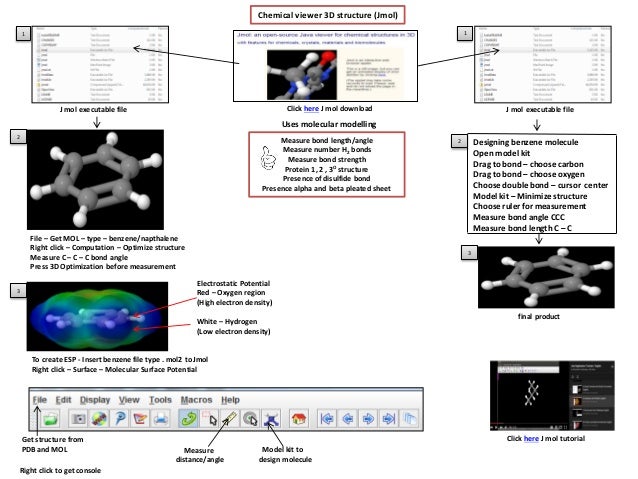
The entire project should first be imported into eclipse. The application is multi-threaded and may be ran from multiple different computers through a network. Folding proteins is an computationally intensive procedure, so we use many simplifications. This application can be used to approximate the 3D structure of proteins. Worked on by Ben Hartshorn, Mike Peters, Travis Reinheimer, Yishan Hu, Chen Yao, Lee Foster, and Aaron Germuth. A Concurrent 3D Protein Folding Application
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
